Technology
Breakthrough DNA Sequencing + AI
The first to industrialize somatic genomics
Beginning in 2022, we set out to build an industrialized somatic genomics company. Now with over three years of somatic data across 15 different diseases, we are operating in a fully scalable manner, incorporating AI and computational biology throughout R&D to maximize our efficiency for our internal and partnered programs.
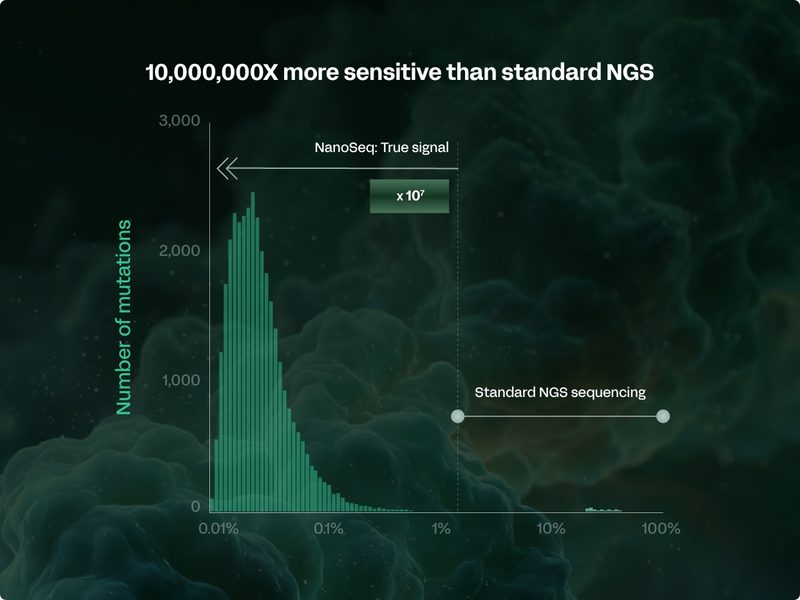
Mining somatic genomes to inform breakthrough medicines
Using the world’s most accurate and sensitive sequencing technology, we uncover the somatic mutations that cause or protect against disease, providing novel, unbiased insights that inform the development of genetically-validated breakthrough medicines.
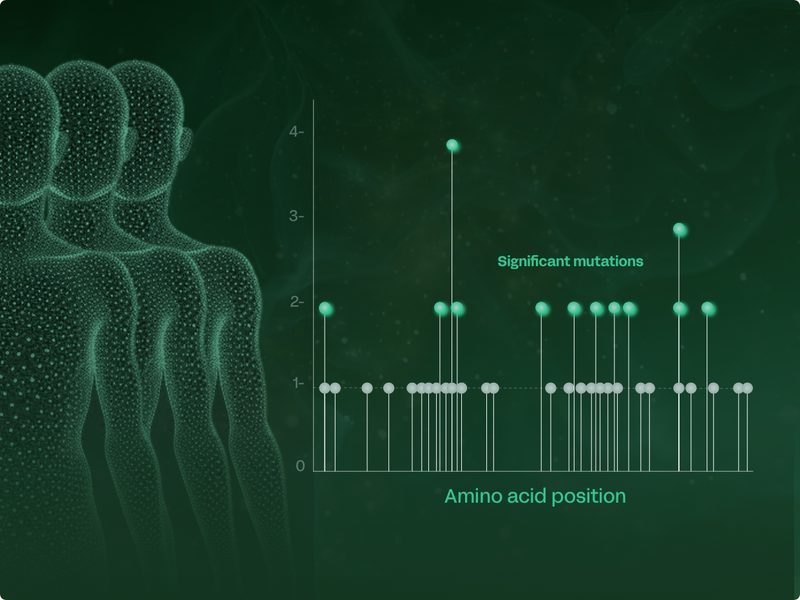
Uncovering disease-causal targets from a handful of patients
By middle age, every cell in our body contains hundreds and sometimes thousands of different mutations. So much so that we find nearly every possible mutation in each organ system. This means that we only need 10 – 30 patients for each study in order to find significant mutations that can drive or protect against disease. In contrast, other genetic approaches require hundreds of thousands of patients to reach significance.
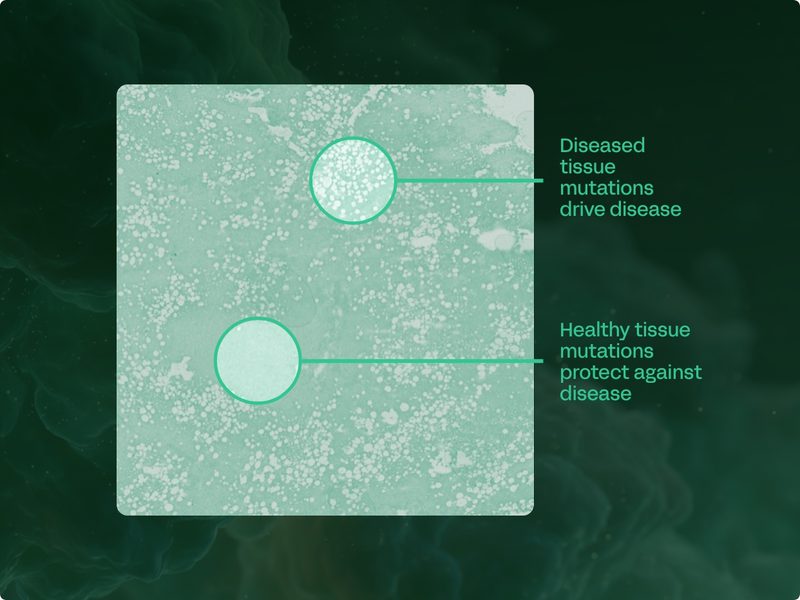
Detecting novel drug targets at the coalface of disease
We detect recurrently mutated genes directly from the cells that are causing or protecting against a given disease. This gives us confidence that the somatic targets we identify are causally implicated in the biology of disease.
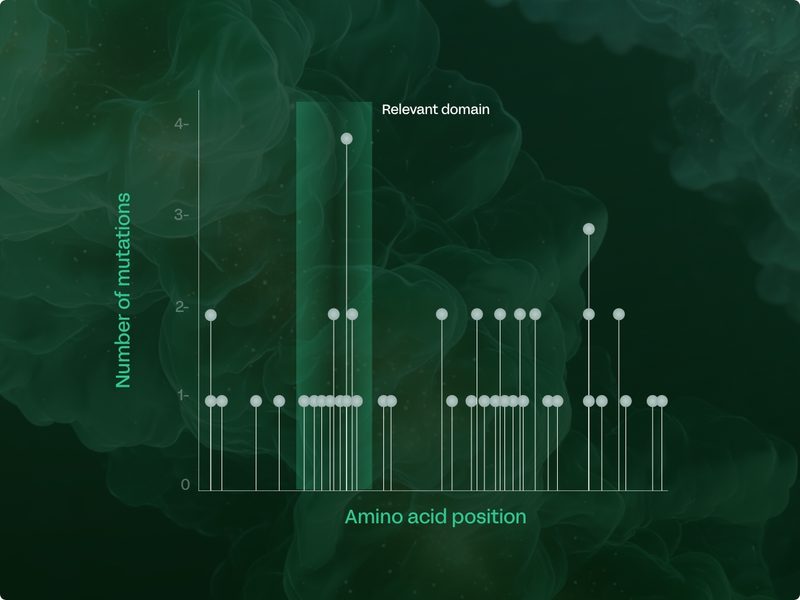
Gaining unprecedented resolution to design better drugs
Somatic mutations not only provide information on disease biology, but also a wealth of information to guide better drug design:
What to target: we often see multiple mutations within a single pathway, allowing for the selection of the most druggable protein within that pathway.
How to target: based on the type of the mutation, the effect of the mutation and the function of the cells in which this mutation is seen, we choose the best therapeutic strategy.
Where to target: analysis of the mutation spectrum along a given gene can highlight specific regions to target for therapeutic effect.
Merging genomics, AI and automation to form a high-throughput, digital biotech
From the moment we receive a patient sample to target identification, biological validation and drug design, we integrate AI and computational tools into our fully digital, data-driven workflows to accelerate our research and development.
Publications
Somatic Genomics has been published in high-impact journals across a broad range of therapeutic areas.
-
2026 Mutations in Time and Space ConferenceApril 22, 2026Drug Discovery in the Age of Somatic GenomicsTim Butler, et al.
-
CellMarch 5, 2026Somatic genomics as a discovery engine for biomedicineSimon F. Brunner, et al.
-
NatureOctober 9, 2025Somatic Mutation Selection at Population ScaleAndrew R.J. Lawson, et al.
-
NatureMay 20, 2021Somatic Mutation Landscapes at Single-Molecule ResolutionFederico Abascal et al.
Somatic Genomics
The Hidden Code of Disease
Pipeline
A breadth of opportunity